Paleontology has for centuries worked to understand the distant past by digging up fossilized remains and analyzing how and why they fit into the evolutionary picture. The results have been impressive.
But they have been limited. The evolutionary picture painted relies largely on the discovery of once hard-bodied organisms, with a smattering of iconic finds of soft-bodied creatures.
In recent years, however, a new approach to understanding the biological evolution of life has evolved under the umbrella discipline of paleogenomics. The emerging field explores ancient life and ancient Earth by focusing on genetic material from ancient organisms preserved in today’s organisms.
These genes can be studied on their own or can be synthetically placed into today’s living organisms to see if, and how, they change behavior.
The goals are ambitious: To help understand both the early evolution and even the origins of life, as well as to provide a base of knowledge about likely characteristics of potential life on other planets or moons.
“What we do is treat DNA as a fossil, a vehicle to travel back in time,” said Betül Kaçar, an assistant professor at the University of Arizona with more than a decade of experience in the field, often sponsored by the NASA Astrobiology Program and the John Templeton Foundation. “We build on modern biology, the existing genes, and use what we know from them to construct a molecular tree of life and come up with the ancestral genes of currently existing proteins.”
And then they ask the question of whether and how the expression of those genes — all important biomolecules generally involved in allowing a cell to operate smoothly — has changed over the eons. It’s a variation on one the basic questions of evolution: If the film of life were replayed from very early days, would it come out the same?

The possibility of such research — of taking what is existing today and reconstructing ancient sequences from it — was first proposed by Emile Zuckerkandl, a biologist known for his work in the 1960s with Linus Pauling on the hypothesis of the “molecular clock.” The molecular clock is figurative term for a technique that uses the mutation rate of biomolecules to deduce the time in prehistory when two or more life forms diverged.
It would not be until decades later that technology and understandings had reached the point where the ideas could become working science. And as Kaçar makes clear, the effort remains a work in progress.
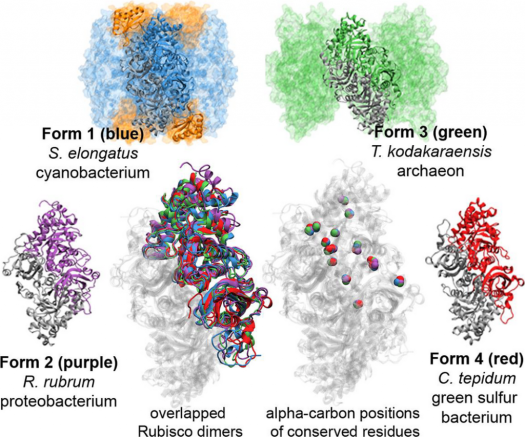
Kaçar is perhaps best known for her work on the RuBisCO enzyme, which is present in plant chloroplasts and is involved in fixing atmospheric carbon dioxide during photosynthesis and producing oxygen. Since life’s origins on Earth, enzymes have been the primary mediators of the chemical reactions inside cells that make life as we know it possible.
This catalyzing of the chemical reaction through which inorganic carbon enters the biological world — processed into sugars and other nutrient-rich compounds — has played an especially central role in the history of life on Earth. RuBisCO is also often described as the most abundant enzyme on the planet.
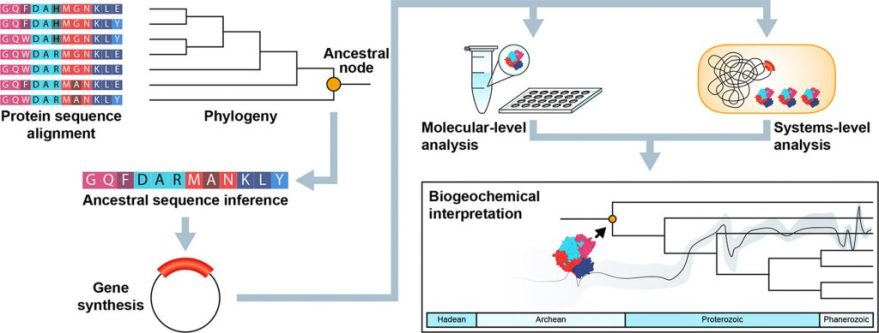
Kaçar and her teams compare RuBisCO and other gene sequences from modern organisms to infer what the sequence must have been in their common ancestor. By doing that many, many times, she says, “we follow back the branches of the evolutionary tree”.
During her time at the NASA Astrobiology Institute as a postdoctoral fellow she developed a system to engineer microbial genomes with synthetic ancient genes, following the path of that inferred RuBisCO tree of life. The protein-producing gene in the modern genomes of an organism such as cyanobacteria can be replaced with an engineered ancestral gene that would have been present perhaps 2.5 billions of years ago, before the rise of oxygen on our planet.
Experiments like this were ongoing when I visited her lab in Tucson earlier this year.
The results: so far, some of the bacteria given the ancestral genes die but some survive while operating more slowly than today’s genes.
That reduced reaction speed — seemingly related not so much to later mutations but to the amount of the enzyme produced by the engineered genome — may have been a feature of the early RuBisCO enzyme, Kaçar said.
The field of paleogenomics, and its search for ancestral forms of important current biomolecules, is growing fast.
“When I first began this work about 10 years ago, I had to painstakingly explain prior foundational ideas by people like Zuckerkandl, just to introduce the basic concept, because people would say, ‘You wanna do what now?'” Kaçar told me.
“Today, people are trying this approach across all different areas of biology and this is great news. For example, I just read about a recent effort to reconstruct and resurrect an ancient ferredoxin–a critical enzyme in shuttling electrons around in metabolism. So the idea is now big enough that others are using the same techniques for their own research interests and questions.”

Kaçar said that her research team has a broad scope. They work on the translation machinery whereby a cell “reads” the information in a messenger RNA and uses it to build a protein; on basic metabolism with enzymes like RuBisCO and nitrogenase, and on biosignatures by looking at photosensitive pigments such as rhodopsons.
“I like that we’re trying to cover many basic biological functions and learning along the way what can or cannot and what ought or ought not be done in developing these techniques.
“We rolled up our sleeves, we have big dreams, the energy and the time. We hope to get there. We are working to get there.”
But the work does have challenges. One is that the ancient rock record biosignatures that Kaçar and her colleagues are looking for — generally different carbon isotopes — may not precisely match the biosignatures left by the enzyme with engineered ancestral genes.
Also, as Kaçar explained, there is little that is “systemically transferable between vastly different ancient molecules.”
“Each mathematical effort to infer what the ancient sequences of a given gene is its own unique undertaking,” she said, “And the model outputs have to be interpreted on their own basis: i.e., is the signal into the deep past good? how far back does the signal reliably go? what is the probability that the inferred sequence will actually be functional?
“In turn, each effort to engineer a genome with an inferred ancient component has to be undertaken on its own merits and may have its own difficulties.”
Kaçar’s work is about as interdisciplinary as could be, bringing together evolutionary biologists and geologists, computer scientists and synthetic biologists, astrobiologists and astronomers. It has also led to her association with, among other organizations, the Earth-Life Science Institute in Tokyo, which focuses on the origins of life.
The implications for origins of life and molecular evolutionary research are pretty clear, but she wants to take the work with ancestral genes and biomolecules further.
“I aim to provide insights into the biology of the past and then tie this knowledge to our search for life in the universe,” she said during a recent Public Library of Science (PLOS) interview.
“A planet free from life today, doesn’t necessary indicate a planet that never hosted life and to understand whether if this was the case, we need to have more than one instance of life to serve as a basis of comparison.
“Earth’s past provides us alternative scenarios. Travel to about 4 billion years ago, what might greet you is a hot, vigorous planet with giant lava volcanoes, no major continents and lots of meteorite impacts.
But we think life, with ecologies different from what is familiar to us today, probably existed back then too.” Surprisingly, however, “molecules of these life forms, however, were close to the fundamental molecules of life today.”
A team at the University of Regensburg in Germany, for instance, found a reconstructed bacteria enzyme they were studying was quite similar to what would have been present 3.4 billion years ago.

Many scientists travel unexpected and intriguing personal paths on the way to their research, and Kaçar is certainly an example of that.
Her family was initially from the Black Sea region of Turkey, where many women are known to be “gözü kara,” which translates from Turkish, Kaçar said, as having a perhaps unwise state of fearlessness. No women in her family had graduated from high school.
Kaçar was born in Istanbul. As she remembers well, while in third grade her father sat her down and told her he couldn’t help anymore with her schoolwork because he had left school after second grade. But he encouraged her to get an education and to become an independent (and fearless) person.

While studying chemistry as an undergraduate at Marmara University in Istanbul, she volunteered to be a real-time translator for a visiting group of scientists who had become to discuss protein biochemistry — especially as it relates to Alzheimer’s and Parkinson’s Diseases.
One talk in particular fascinated her. It was on the molecular properties of an enzyme whose activity changes with age. She says she will never forget that first time she learned that when the site of a protein changes, entire cellular and bodily activities can collapse.
She was “hooked” and reached out to one of the American professors who had been in the group. He suggested that she apply for a Howard Hughes Medical Institute summer undergraduate research scholarship, which she did and won one. That brought her to Atlanta and Emory University, where she visited his laboratory to study a particular protein . This was her first time in the U.S. or in a scientific research laboratory. In 2004, I came back to the same laboratory as a graduate student. She was 20 years old and didn’t know anyone outside the lab.
In the following years she won a NASA research grant, taught and researched at Harvard University, won her current position at UA and, last year gave birth to a son. Kaçar is an accomplished public speaker as well, dedicated to bringing science to people.
And all the while, she is working to solve major scientific problems in a burgeoning but exacting discipline.
“I am not afraid of failure,” she said. “For better or for worse, I am “gözü kara.”‘
